Giann Gorospe, Renjun Zhu, Michal A. Millrod, Elias T. Zambidis, Leslie Tung, Rene Vidal. Johns Hopkins University, Volume 61, Issue 9, Page:2389-2396
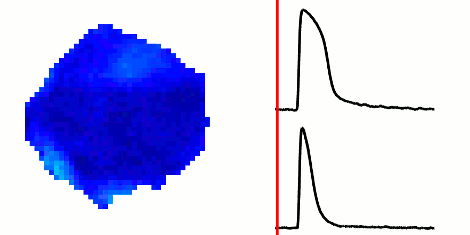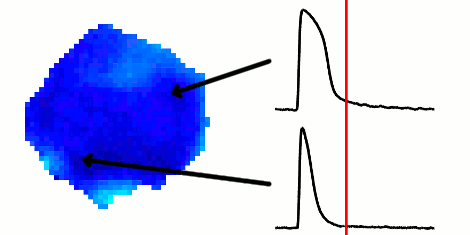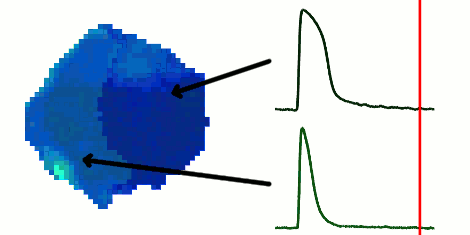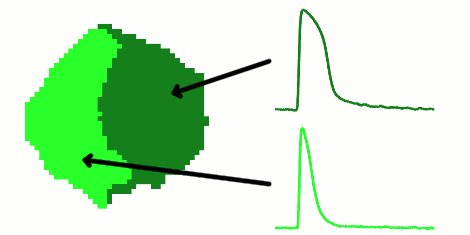
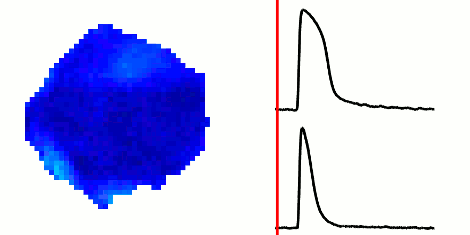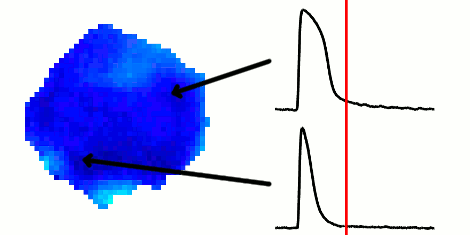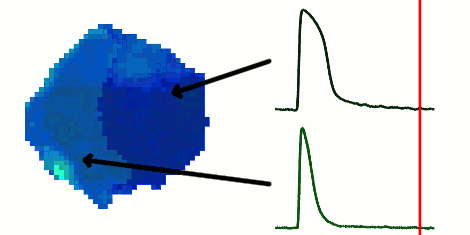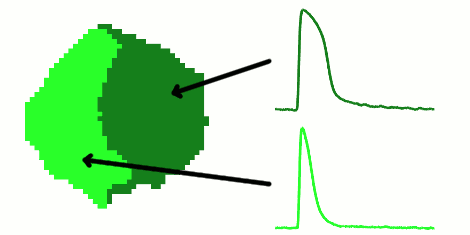
The rapid advancement of stem cell research in all fields of medicine over the last decade has been outstanding. However, in the area of cardiac regeneration where cardiomyocytes are differentiated from stem cells, our general understanding of the de novo cell population is still far from adequate. Clinical applications of these cells require systematic methods for determining phenotypes of cardiomyocytes (such as ventricular-like, atrial-like, nodal-like), which are not yet well established. In the absence of reliable molecular markers that can be used on live cells, a way to approach this problem is from an electrophysiological perspective, using the action potential. Current methods for phenotype identification apply ad hoc criteria to action potential features such as duration and amplitude of the action potential. Automated, full signal based approaches to grouping cardiomyocytes may prove to be more effective, since these approaches remove the subjective nature of manual segmentation, and allow for scalability to large datasets of cardiomyocytes as well as consistency in discrimination across research groups. In this paper, we use optical mapping to obtain action potential recordings from differentiated cardiomyocytes in embryoid bodies, and apply methods from signal processing and machine learning to perform automated grouping of the cardiomyocyte population based on their action potential waveforms. These methods allow us to assess population heterogeneity, and in the future can help to refine culture conditions to enhance desired cellular phenotypes.
Keywords: Cardiac Electrophysiology, Cardiomyocyte, Action Potential, Optical Mapping, Spectral Grouping, Signal Processing