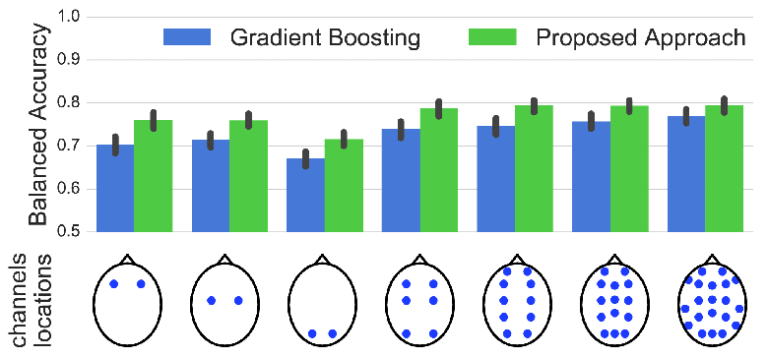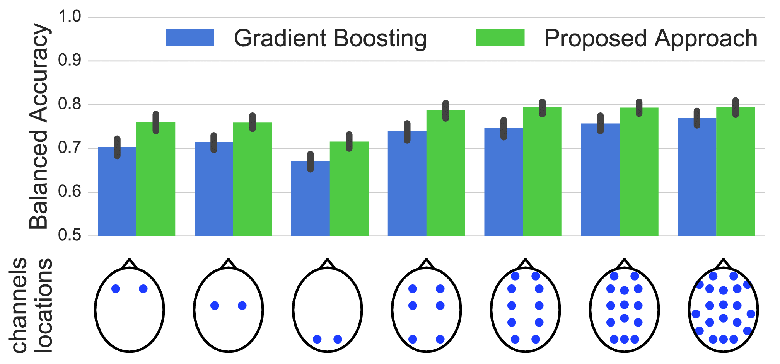
Sleep stage classification constitutes an important preliminary exam in the diagnosis of sleep disorders. It is traditionally performed by a sleep expert who assigns to each 30 s of the signal of a sleep stage, based on the visual inspection of signals such as electroencephalograms (EEGs), electrooculograms (EOGs), electrocardiograms, and electromyograms (EMGs). We introduce here the first deep learning approach for sleep stage classification that learns end-to-end without computing spectrograms or extracting handcrafted features, that exploits all multivariate and multimodal polysomnography (PSG) signals (EEG, EMG, and EOG), and that can exploit the temporal context of each 30-s window of data. For each modality, the first layer learns linear spatial filters that exploit the array of sensors to increase the signal-to-noise ratio, and the last layer feeds the learnt representation to a softmax classifier. Our model is compared to alternative automatic approaches based on convolutional networks or decisions trees. Results obtained on 61 publicly available PSG records with up to 20 EEG channels demonstrate that our network architecture yields the state-of-the-art performance. Our study reveals a number of insights on the spatiotemporal distribution of the signal of interest: a good tradeoff for optimal classification performance measured with balanced accuracy is to use 6 EEG with 2 EOG (left and right) and 3 EMG chin channels. Also exploiting 1 min of data before and after each data segment offers the strongest improvement when a limited number of channels are available. As sleep experts, our system exploits the multivariate and multimodal nature of PSG signals in order to deliver the state-of-the-art classification performance with a small computational cost.