Advanced CRISPR technology is part of a new test that determines within minutes whether a person has been infected with even very low levels of the SARS-CoV-2 virus (the cause of COVID-19), and also quickly and directly measures the viral load, or how much replicating virus is in a person’s body [1].
The California research group behind the technology seeks to provide a do-it-yourself test that people can use to check themselves for infection before going out in public, so they know whether they should head to work or school, hop on a plane, or visit older relatives. If an infection is found, clinicians can also use the test’s report of viral load for such purposes as making a prognosis, determining how contagious a person is, and tracking patient progress to inform treatment.
A few CRISPR-based tests have already become available, but they do not directly measure viral load. This is because they rely on amplification of deoxyribonucleic acid (DNA) as a first step. Since SARS-CoV-2 has ribonucleic acid (RNA) as its genetic material, these tests must convert the viral RNA into DNA. Then they amplify the DNA via polymerase chain reaction (PCR) or other amplification methods to make the presence of the starting RNA more obvious and finally detect the resulting DNA using CRISPR enzymes or conventional DNA measurements, explained Daniel Fletcher, Ph.D., professor of bioengineering and biophysics at the University of California, Berkeley, and part of the California research team that developed the new COVID test. This multistep process is good at determining whether the virus is present, but it can obscure information about the patient’s original infection, he said. “At some point you lose your understanding of precisely how many copies of viral RNA you started with. Was it one, was it 10, was it 1000?”
One of the keys to the California group’s new technology is an enzyme called Cas13, especially one known as Cas13a [2]. Cas13a works on RNA and therefore skips the RNA-to-DNA-conversion. The RNA-detecting ability of the CRISPR Cas13 enzyme [2] was first identified by another member of the California research group: Jennifer Doudna, who is the Li Ka Shing Chancellor’s Professor of Biomedical Science at UC Berkeley, and also a winner of the 2020 Nobel Prize in Chemistry for her groundbreaking CRISPR research.
With Cas13’s application for RNA detection, it has the potential to diagnose infection with the novel coronavirus and also report the patient’s viral load. The challenge was whether there would be enough signal to make the measurement without amplification.
Targeting RNA
To get their new approach to work, the California research group had to provide a way for Cas13 to find its target, SARS-CoV-2, and then go about its primary task, which is to cleave the viral RNA. The researchers accomplished this task by developing multiple guides. Each guide was a short RNA sequence designed to home in on different identifying sections of the SARS-CoV-2 genome (Figure 1). The use of multiple guides should help to ensure Cas13 would locate the target and generate a stronger signal at the same time.

That strategy worked. In fact, multiple guides allowed Cas13 to detect as few as 30 copies of target RNA per microliter, and since infectivity of SARS-Co-V-2 typically starts at around 1000 copies per microliter, this was a key finding, said group member Melanie Ott, M.D., Ph.D., director of the Gladstone Institute of Virology and professor of medicine at the University of California, San Francisco (Figure 2). “The interesting part of the project began when we developed the guides and saw that by combining two guides together, we could actually push sensitivity into a direction that was diagnostically interesting for the coronavirus. Once we realized that there was a possibility there, we all became very active in pursuing this.”

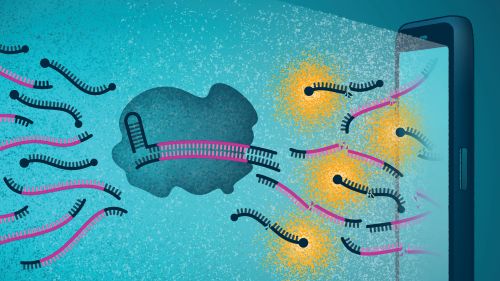
Beyond detecting SARS-Co-V-2, the researchers also wanted to measure how much virus was present. For that, they introduced fluorescence to the mix. Specifically, they used fluorescent probes that only emit their fluorescence when Cas13a has recognized the viral RNA and cleaved a so-called quencher molecule from the probe (Figure 3). Consequently, Ott explained, the higher the concentration of viral RNA, the more Cas13 cleaving activity, and more cleaving activity translates to a greater accumulation of fluorophore. In other words, the rate of fluorescence emission reveals the viral load. She added, “This is the uniqueness of the assay. It gives us a direct quantitative assay without reverse transcription and amplification.”
Bringing in the smartphone
As the researchers began to realize they could get a reliable and quantifiable signal of Cas13 activity through fluorescence, they began developing a convenient and quick way to access that measurement.
“Since we weren’t making more copies of the viral RNA through amplification and were instead simply acting on the existing viral copies, that presented us with an opportunity to measure how many viral copies are present, potentially even outside of a standard clinical laboratory,” Fletcher said (Figure 4). That is where the smartphone came in.

Fletcher and his group got to work developing what he described as “essentially a compact fluorescence microscope.” The answer was literally at their fingertips. “Thanks to consumers buying lots and lots of mobile phones, and to phone companies making their cameras better and better to attract more customers, this cycle has led to extremely high-quality cameras that are suitable for sensitive fluorescence detection,” he said. “And by using a phone to prototype a diagnostic device, we can simplify a range of other steps: the user interface is built into the phone, and so are the processing power to analyze those images, and the ability to control external components, such as turning on and off a light source to illuminate the sample. The phone ends up being a very powerful control center, in addition to providing a camera for detection.”
Pulling it all together
The next step is to package the technology into an easy-to-use, fully automated test kit: the goal is to have users run a swab shallowly around the inside of the nose, pop the swab into a cartridge, and press a start button on their phone, Fletcher described. The researchers are already entrenched in the development of the kit (Figure 5). He estimated they would need several months to optimize it and then apply to obtain the necessary U.S. Food and Drug Administration authorization. In the meantime, they are in preliminary discussions to find a manufacturer.

(Image courtesy of UC Berkeley.)
While that proceeds, the research group is also optimistic about the technology’s potential for detecting other infections. “That’s the beauty of CRISPR and this platform. As long as it’s still carrying RNA and is respiratory captured by a nasal swab, all that is necessary is to adapt the guide sequence to new pathogens,” Ott said. Such a test could help a person determine whether they had a common cold or influenza, or could possibly distinguish different strains of the COVID-19 virus, she remarked.
Fletcher hopes this technology and others ongoing in the research community will help improve medical care overall after the pandemic. “The current crisis has really revealed how inadequate our health care system is in terms of detecting infectious diseases like viruses. For instance, we are asking patients all to come to a central location for testing, which is pretty much the worst way to do this when we’re dealing with something that’s communicable,” he remarked. “For all the tragedies surrounding this pandemic, I think it has also given us this motivation and this opportunity to push for new technologies and new approaches to diagnosis.”
Ott agreed. “I believe this past year will not just be a lost time. We are being inventive, we are trying new approaches, and I think good things will come out of it,” she said. “I hope this drive to find solutions will continue, because we haven’t heard the last of this. The next pandemic will come. We need to be ready.”
References
- P. Fozouni et al., “Amplification-free detection of SARS-CoV-2 with CRISPR-Cas13a and mobile phone microscopy,” Cell, vol. 184, no. 2, pp. 323–333.e9, Jan. 2021, doi: 10.1016/j.cell.2020.12.001.
- A. East-Seletsky et al., “Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection,” Nature, vol. 538, no. 7624, pp. 270–273, Oct. 2016.



