Gene editing and CRISPR (a group of repeated DNA sequences in bacteria) typically target disease-causing mutated genes by eliminating the bad gene altogether, by correcting the problem DNA to restore proper gene functioning, or by modifying a different gene to compensate for the faulty gene’s lost function. One research group at Duke University in Durham, North Carolina, however, is using a different strategy to fight one of the most common inherited genetic diseases: Duchenne muscular dystrophy (DMD).
DMD is a severe degenerative muscle disorder that often arises from a frameshift mutation caused by the deletion of a region of the gene that throws off the reading of the gene’s instructions and, consequently, impedes its ability to produce its protein. In this case, the protein is dystrophin, a key part of the dystrophin-glycoprotein complex that forms a long chain between the cytoskeleton and the cell membrane in a muscle cell, holding that cell together when force is applied, explains Charles Gersbach, Rooney Family Associate Professor of Biomedical Engineering at Duke University (below).

Gersbach and his group are taking advantage of the fact that most DMD mutations occur in the midst of multiple sequence repeats located at the middle of the gene; so, rather than having to correct the mutation itself, the researchers can remove just enough of the surrounding DNA to get the reading frame back on track [1]. “The repeated sequences essentially produce links within this long chain, and we have learned from another disease, called Becker muscular dystrophy, that you can actually remove links from that chain, and, in many cases, the protein remains largely functional,” he says. “For that reason, we don’t have to restore the gene back to the normal wild-type sequence, and instead we can cut out exons to restore the reading frame.”
Cutting out DNA is a good option for several reasons, Gersbach notes. One is that muscle cells are no longer dividing “so they basically shut down the pathways that allow you to change or to replace DNA. That means, to get sufficiently high efficiencies, you really are stuck with just removing segments of DNA.” Another reason stems from the diversity of mutations that cause DMD. “Every patient has a different mutation, so if you wanted to change each individual patient’s mutation back to the wild-type sequence, you’d have to have a custom gene therapy for every single one of them,” Gersbach explains, adding that no obvious path exists for clinical trials of customized gene therapies.
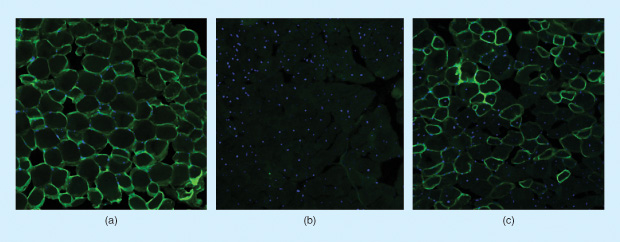
His group’s approach, on the other hand, is a single gene-editing mechanism that can capture a large group of DMD patients. So far, the Duke researchers have shown that the technique is effective in resetting the reading frame in patient-derived muscle cells and in changing the outcome of the disease in mouse models (below). They have published work showing that the removal of one exon can address 13% of the entire DMD patient population, whereas the removal of a larger region spanning 11 exons will cover about half of the patient population. “Although we haven’t done all the variations,” Gersbach continues, “you can imagine a number of different ways in which you could get to almost all of the DMD patients.”
The Duke researchers are delivering CRISPR to muscle tissues via a version of adenoassociated virus (AAV), which enters the bloodstream and reaches all of the muscle tissue throughout the body. Although this method allows for systemic distribution, it is not perfect. “For one thing, it isn’t absolutely specific to muscle, so it does go to some other tissues as well,” Gersbach reports. Perhaps the biggest problem, however, is that AAV has a tendency to linger. “With CRISPR, you want more of a hit-and-run approach, where you go in, edit, fix the mutation, and then you want the CRISPR system to go away. But with AAV, it’s possible that the CRISPR system is going to stick around for a while. We’re not sure if that will happen, but that is a concern because it might end up cutting places in the genome that it shouldn’t.”
Currently, the group is optimizing the vector system, generating new mouse models that have the human gene, and working with collaborators on larger animal models. The research is still early in development and faces some obstacles, Gersbach concedes, but it is a worthy project. He remarks, “Muscular dystrophy is definitely not a low-hanging fruit for gene editing, but it’s potentially one of the more impactful because there are no other options for a lot of these patients and this could help. That’s exciting.”
References
- J. N. Robinson-Hamm and C. A. Gersbach, “Gene therapies that restore dystrophin expression for the treatment of Duchenne muscular dystrophy,” Human Genetics, vol. 135, no. 9, pp. 1029–1040, Sept. 2016.



