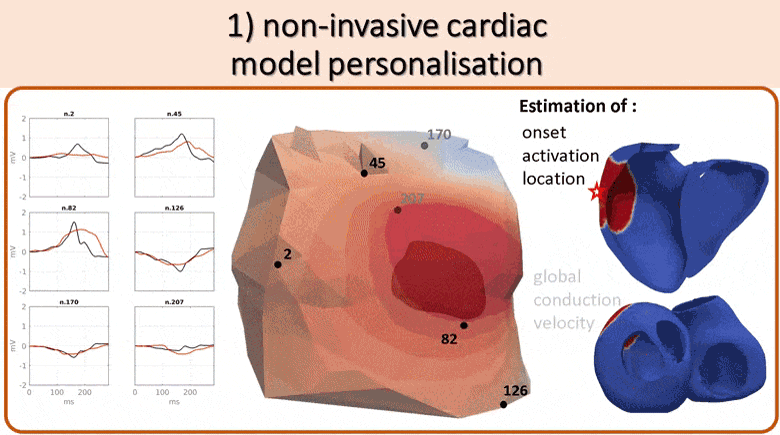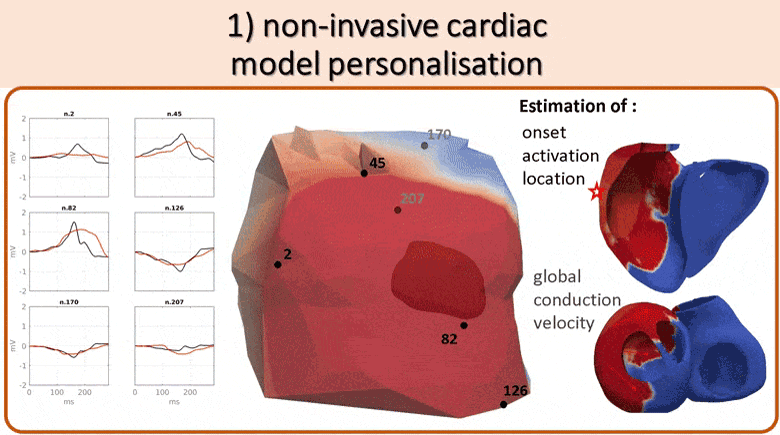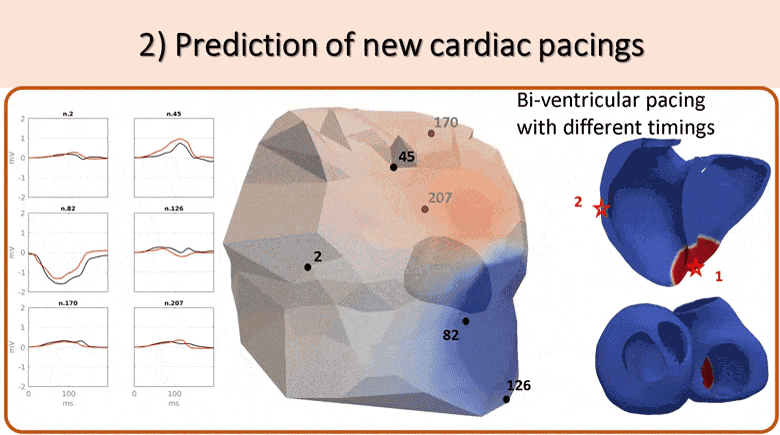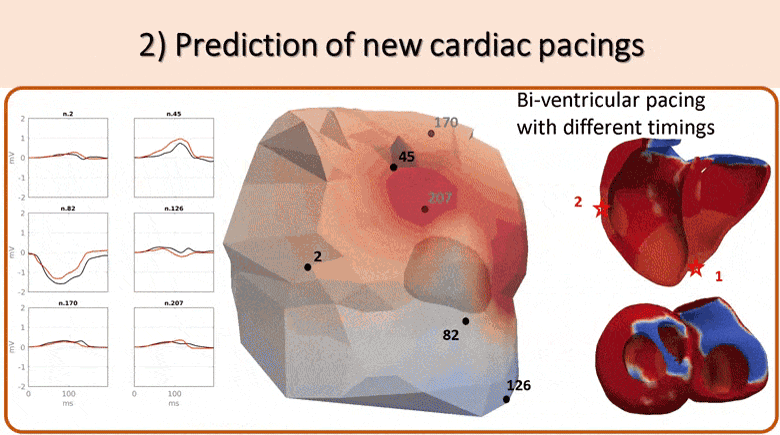
We use non-invasive data (body surface potential mapping, BSPM) to personalize the main parameters of a cardiac electrophysiological model for predicting the response to different pacing conditions.
First, an efficient forward model is proposed, coupling the Mitchell-Schaeffer transmembrane potential model with a current dipole formulation. Then we estimate the main parameters of the cardiac model: activation onset location and tissue conductivity. A large patient-specific database of simulated BSPM is generated, from which specific features are extracted to train a machine learning algorithm. The activation onset location is computed from a Kernel Ridge Regression and a second regression calibrates the global ventricular conductivity. The evaluation of the results is done both on a benchmark dataset of a patient with premature ventricular contraction (PVC) and on 5 non-ischaemic implanted cardiac resynchonisation therapy (CRT) patients with a total of 21 different pacing conditions. Good personalisation results were found in terms of the activation onset location for the PVC (mean distance error, MDE=20.3mm), for the pacing sites (MDE=21.7mm) and for the CRT patients (MDE=24.6mm). We tested the predictive power of the personalized model for biventricular pacing and showed that we could predict the new electrical activity patterns with a good accuracy in terms of BSPM signals. This is an encouraging first step towards a non-invasive pre-operative prediction of the response to different pacing conditions to assist clinicians for CRT patient selection and therapy planning.

