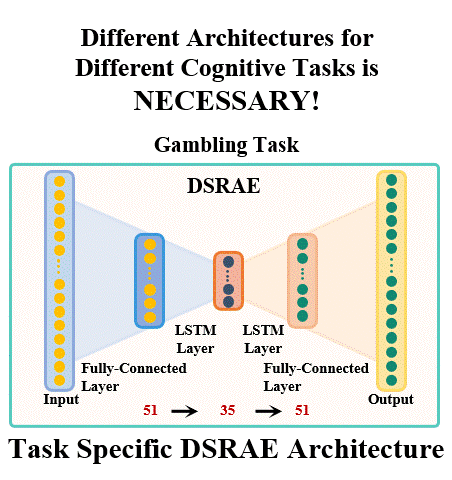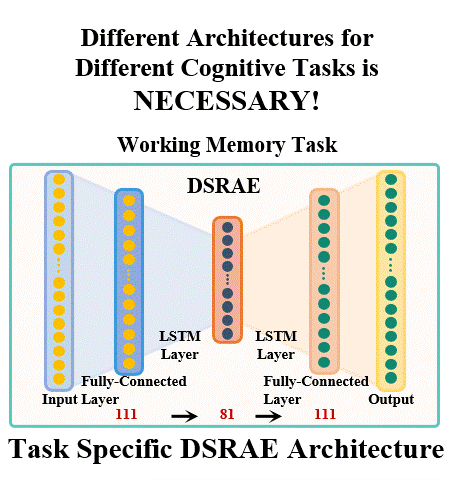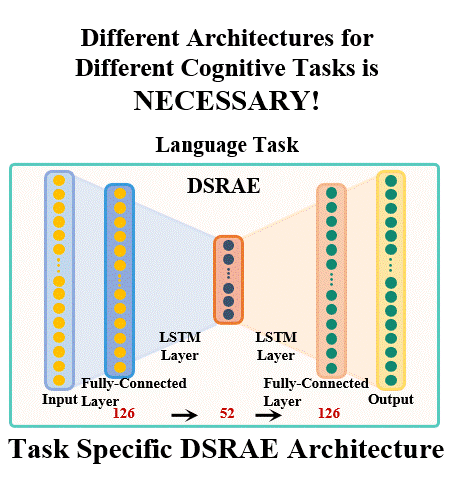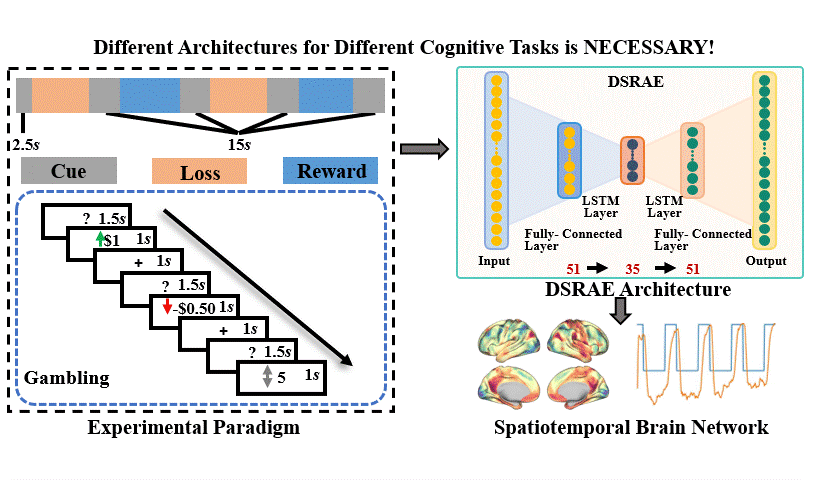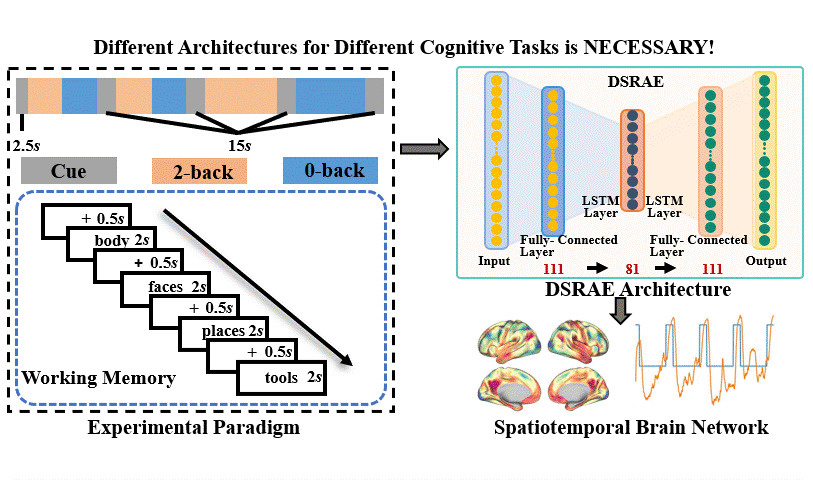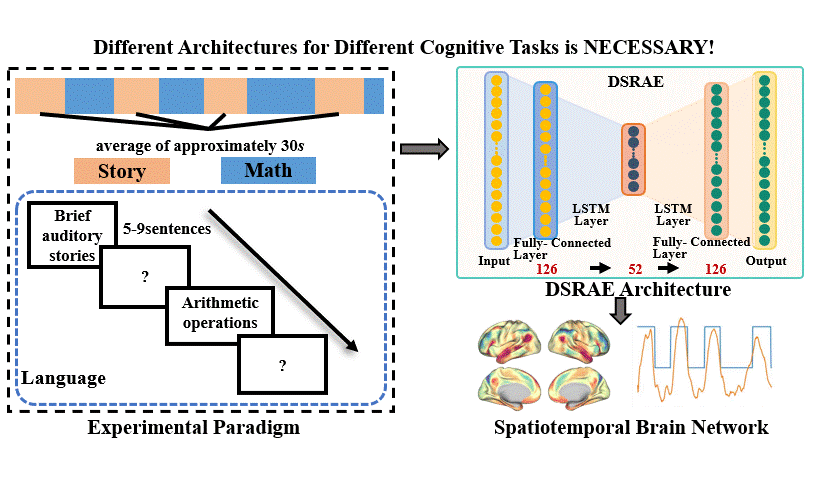
The brain undergoes complex neural processes that are highly correlated spatially and temporally. Using deep neural networks (DNNs) to explore the spatial patterns and temporal dynamics of human brain activities has been an important yet challenging problem because the artificial neural networks are hard to be designed manually. There have been several promising deep learning methods, e.g., deep sparse recurrent auto-encoder (DSRAE) that can decompose neuroscientific and meaningful spatiotemporal patterns from 4D functional Magnetic Resonance Imaging (fMRI) data. However, the previous studies still depend on hand-crafted architectures and hyperparameters, which are not optimal in various senses, i.e., across different cognitive tasks. That would affect the performance and efficiency of spatiotemporal brain network decomposition for the specific tasks. In this way, we aim to propose a novel framework to optimize such DNN model for spatiotemporal brain network decomposition. We employ the light-weighted evolutionary algorithms (EA) to optimize the architecture of DSRAE by minimizing the expected loss of initialized models, named eNAS-DSRAE (i.e., evolutionary Neural Architecture Search on Deep Sparse Recurrent Auto-Encoder). Validation experiments are designed and performed on the publicly available human connectome project (HCP) 900 datasets, and the results achieved by the optimized eNAS-DSRAE suggested that our framework can successfully identify the spatiotemporal features and perform better than the hand-crafted neural network models and classical traditional brain network analysis methods. To our best knowledge, the proposed eNAS-DSRAE is not only among the earliest NAS models that can extract meaningful spatiotemporal connectome-scale brain networks from 4D fMRI data, but also is an effective framework to optimize the RNN-based models.