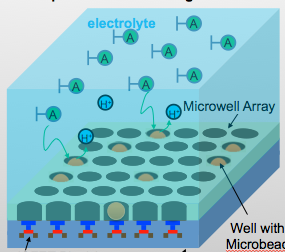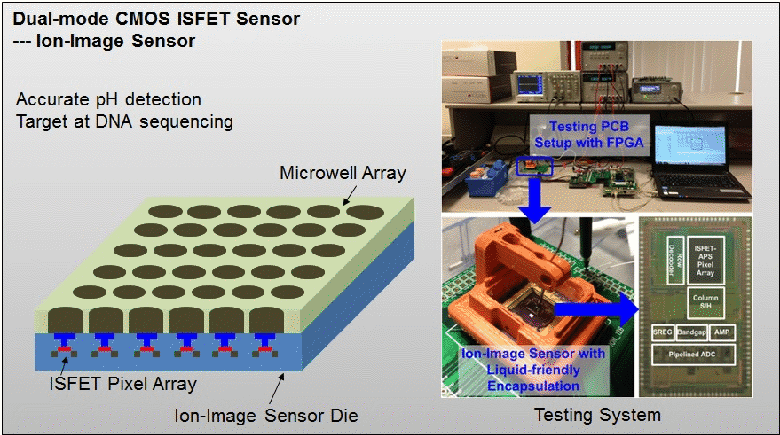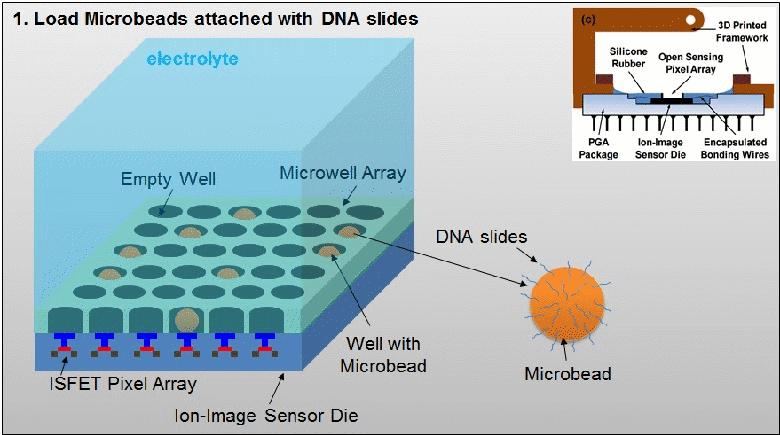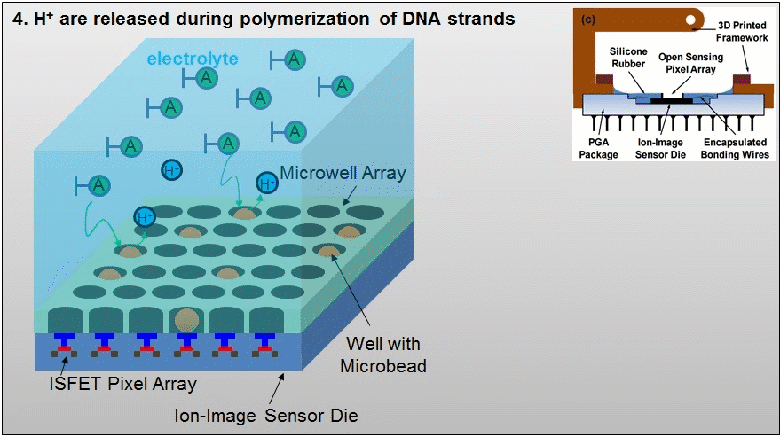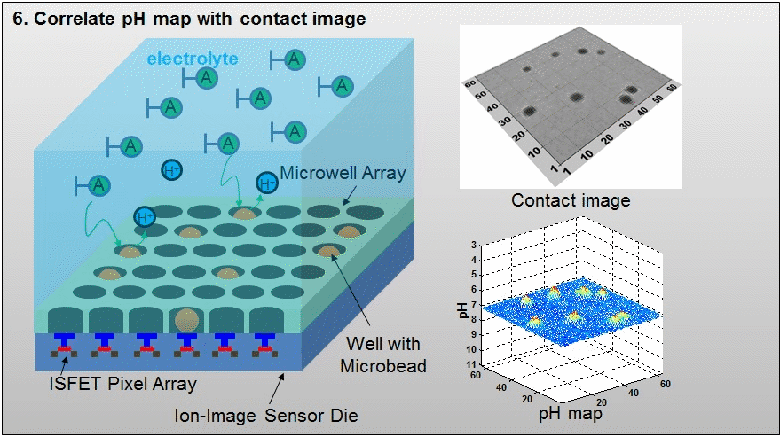
The existing ion-sensitive field effect transistor (ISFET) based DNA sequencing detects hydrogen ions (H+) released during polymerization of DNA strands that are attached on microbeads. However, these microbeads are scattered into microwell array above the ISFET sensor with unknown distribution. As such, false pH detection happens at empty microwells in traditional ISFET-based DNA sequencing due to cross-talk from neighbouring microwells. In this paper, a dual-mode CMOS ISFET sensor is proposed to have accurate pH detection towards DNA sequencing. The dual-mode sensing, namely optical and chemical modes, is realized by integrating CMOS image sensor (CIS) with ISFET pH sensor, which is fabricated in standard 0.18 μm CIS process. With accurate determination of microbead physical locations with CIS pixels by contact imaging, the dual-mode sensor can correlate local pH for one DNA slice attached at one location-determined microbead. This can result in significantly improved pH detection accuracy. Moreover, towards high-throughput DNA sequencing, a correlated double sampling (CDS) readout circuit that supports large dual-mode sensor array is deployed to reduce pixel-to-pixel non-uniformity such as threshold voltage mismatch. The proposed CMOS dual-mode sensor is experimentally examined to show a well correlated pH map based on the contact-imaging of microbeads. The state-of-the-art measured results show: a pH sensitivity of 26.2mV/pH, a fixed pattern noise (FPN) reduction from 4% to 0.3%, and a readout speed of 1200 frames/second (fps). Therefore, the developed dual-mode CMOS ISFET sensor has great potential for future personal genome diagnostics with high accuracy and low cost. For more information, please visit our group website at www.ntucmosetgp.net.