From “ridiculous” DARPA challenge to COVID-19 prevention and treatment
Long before the coronavirus-2019 (COVID-19) pandemic began, four research groups—two at universities and two at biotechnology companies—were preparing for it by tackling a seemingly outlandish challenge proffered by the Defense Advanced Research Projects Agency (DARPA) of the U.S. Department of Defense: find a stopgap defense against emerging pathogens, so people would have protection until a longer-term solution, notably a vaccine, became available. Under the challenge, called the Pandemic Prevention Platform (P3) program, the groups were to develop technology that would generate a treatment within 60 days of a virus being identified, and have that treatment begin to confer protection within three days of it being administered [1].
“When the P3 program was rolled out, everyone obviously thought it was ridiculous because the typical fastest drug programs are in the 2- to 5-year range. So to think about doing something in two months didn’t even seem plausible,” recalled James Crowe, Jr., M.D., director of the Vanderbilt Vaccine Center at Vanderbilt University in Nashville, TN (Figure 1). Nonetheless, four groups felt the premise was so interesting—and important— that they took up the gauntlet. They included the Crowe-led research team at the Vanderbilt Vaccine Center; a research group at the Duke Human Vaccine Institute at Duke University in Durham, NC; and teams at the company AbCellera Biologics Inc. in Vancouver, BC, Canada, and at the AstraZeneca subsidiary MedImmune of Gaithersburg, MD.

By late 2019, all four had made considerable headway developing antibodies that could be used as short-term treatment for, as well as prevention against, various model viruses. Unlike longer-lasting vaccines that train the body to recognize viruses so it can mount an immune/antibody response—a process that typically takes at least a couple of weeks—the P3 antibody approaches required no immune-system training. Rather, the research groups either used lab-produced, protein delivered antibodies that could be introduced to the body as a nearly instantaneous defense, or antibody- making instructions delivered via a snippet of mRNA code so the body’s immune system could start churning out antibodies within days.
In the midst of that work, COVID-19 struck, and each of the groups took what they had already learned and redirected their attention to the real-life viral threat that was spreading around the world: severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Today, protein-derived antibody therapies are now in trials and could be ready for patient use by early 2021, and work on ribonucleic acid (RNA) delivered antibody therapies is well under way and might make their way to the clinic by early 2022.
mRNA-delivered antibodies
For the P3 program, the Duke Vaccine Institute research group had been studying a variety of model organisms in 2018 and 2019, including a respiratory virus, specifically influenza. “We had gotten to the proof-of concept point in November/December of 2019, and then wham! We had the potential to help with COVID-19 and we started racing,” described Thomas Denny, chief operating officer of the Duke Human Vaccine Institute and professor of medicine at Duke University. DARPA encouraged the Duke grou to pursue its approach, which was mRNA delivery of antibodies, and in June 2020 provided the group with additional funding to get the method ready for phase 1 clinical trials by the end of 2020 or early 2021.
The Duke team is pursuing the delivery of exogenous mRNA that encodes for COVID virus-neutralizing antibodies [2]. The recipient’s cells transcribe the mRNA and begin expressing the antibodies as directed, explained Gregory Sempowski, Ph.D., who is leading the Duke P3 effort, and is also professor of medicine and pathology at Duke’s School of Medicine (Figure 2). “The antibodies then travel around the body and give passive protection for a short period of time,” he said, noting that he expects protection to last for about 30 days, after which subsequent doses could potentially extend the protection.

The decision to pursue mRNA as the avenue to fight SARS-CoV-2 was made easier because the Duke Human Vaccine Institute had already been developing mRNA-delivered vaccines against human immunodeficiency virus (HIV) and influenza for a number of years, “so it was a natural transition to take that platform technology for delivering immunogens, or the recombinant protein components of a vaccine, and just pivot it over to delivering the actual antibody,” Sempowski said.
Using blood samples from infected individuals, the researchers identified B cells that were making the SARS-CoV-2 antibodies, he said. “Once we did, we could sort out those individual cells, clone their sequences, and in a matter of six weeks, we were able to generate 2500 potential mRNA sequences [for making potent SARS-CoV-2 antibodies],” Sempowski said. After that, down-selection began. This involved expressing the antibodies from those sequences, producing the antibodies on a small scale, and running screening and functional assays in specialized biosafety level-3 facilities at Duke to figure out which sequences yielded antibodies that could successfully neutralize the SARS-CoV-2 virus (Figure 3).
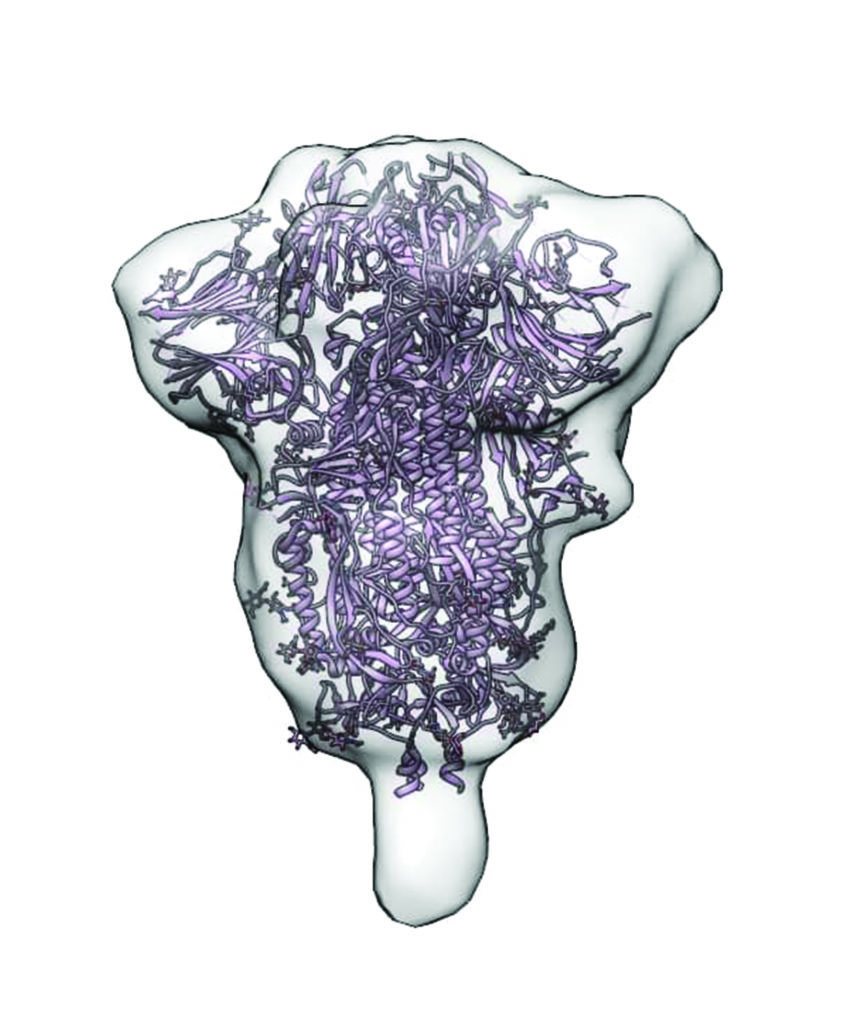
In particular, the researchers are focusing on those sequences that produce antibodies capable of binding to a structure on the virus’s membrane, called the spike protein (Figure 4). Such binding disables the spike protein and prevents the virus from entering human cells. “We are now mixing these very strong candidate sequences with the live virus to verify that the live virus gets inhibited,” Sempowski said.

Based on those results, they have focused on one or two especially attractive contenders to begin running tandem animal-challenge models: one that directly administers antibodies to verify that they are effective against the virus, and a second that administers mRNA encapsulated in a lipid nanoparticle carrier to ensure that the animal can use it to make the antibodies on its own.
The Duke team has partnered with the research group of Drew Weissman, M.D., Ph.D., professor in the Perelman School of Medicine at the University of Pennsylvania, to identify the optimal lipid-nanoparticle carrier to transport the mRNA safely to the cells after it is administered via small-volume (likely intravenous) infusion. “There are some nuances to this, so we’ve spent a lot of time working with Drew’s group to design mRNA-expression constructs, and to identify and down-select the optimal lipid formulation for the maximal delivery and expression of the antibody in animal models,” Sempowski said.
He anticipates the mRNA approach go into phase 1 human trials by the beginning of 2021 with clinical availability about 12 months afterward. Sempowski acknowledges that this timeframe is longer than antibodies in protein format (detailed below) because it is a new methodology that will require additional scrutiny from the U.S. Food and Drug Administration (FDA) to prove that it is safe, causes no adverse reactions, and achieves good levels of protection. “There’s only been one other mRNAdelivered antibody [3] and that was a chikungunya antibody from Moderna (in concert with Vanderbilt University),” he said, “so every time we do these types of studies, it adds more to the profile of the feasibility of this new platform.”
Antibodies in protein format
Protein-delivered SARS-CoV-2 antibodies are further along in their development, and several are already in clinical trials, including antibodies made by the Vanderbilt research group. Crowe anticipates both will provide a patient with protection for about three months.
At Vanderbilt, the group set out on the DARPA P3 challenge with a simulated outbreak, but used Zika as its model instead of influenza. “We took a blood sample from someone who had been infected, clicked a stopwatch and said go, and then we went as fast as we could to make human monoclonal antibodies, and we went from blood sample through discovery mice and to a nonhuman primate protection study in 78 days. That was really the fastest in history that anyone had done that,” Crowe said, noting that the studies are described in detail in [4] and [5].
The Vanderbilt researchers were preparing to embark on a second outbreak simulation in January, when “on a dime, we shifted from thinking about simulating a bird flu outbreak to doing the real thing with coronavirus,” Crowe said. By the third week of the month, they had obtained a blood sample from the first U.S. case in Seattle “while the person was still in the hospital and infected,” and using the genetic sequence of SARS-CoV-2, which had just been published by Chinese scientists, they then synthesized the viral DNA and made proteins to represent the virus, he described. “That gave us protein for binding assays, and the virus for functional assays, and that was all we really needed for our discovery program.”
In all, the group isolated and characterized about 2000 antibodies and down-selected to about 30 that were of the highest potency against SARSCoV- 2. “From the time we got the blood sample to the time we handed over monoclonal antibodies (identical antibodies lab-generated from a single B-cell clone) to our commercial partners for clinical development was 24 days. It was just amazing,” Crowe said. AstraZeneca is now proceeding with two of them for an antibody cocktail, and the startup IDBiologics of Nashville, TN, is moving forward with one in a monotherapy. He said each approach has its benefits: two doses makes doubly sure that one antibody or the other will stop the virus; whereas a monotherapy requires a more straightforward and consequently a likely faster clinical-trial process and path to FDA approval, and also has a simpler manufacturing process, which should generate more doses. “There is not a fixed opinion about what’s best,” he remarked.
Whether its monotherapy or a cocktail, manufacturing may be an issue, he added. “There is concern (from public health authorities) about whether there’s going to be enough capacity to make what is needed. The manufacturers are saying they can make up to a million doses, which sounds like a lot, but there are 300 million people in the U.S., so you wouldn’t even have enough for 1 percent of the population.” He anticipates a flurry of engineering innovation to arise over the coming months to help solve that bottleneck.
In the meantime, the Vanderbilt group is continuing to investigate RNA-delivered antibodies, which have reduced manufacturing demand, and has also begun focusing on the next viral pathogen that comes along, because one will, Crowe asserted. “We’ve had (avian influenzas) H5 and H7, Zika, chikungunya, Ebola—it’s just like clockwork.” To that end, he and his research group began a program called AHEAD100, for which they have assembled a list of known and potentially dangerous viruses and are methodically making monoclonal antibodies to all of them so they have them ready to go when the call comes (Figure 5). “We are already about 40 viruses into this, so we have antibodies for North American and South American hantaviruses, Rift Valley fever virus in East Africa, Ross River virus in Australia, and others you’ve never heard of, because we know they’re in mosquitoes, bats, or other (vectors) from which they can cross over into humans.”
Forward-thinking efforts, such as the DARPA P3 or AHEAD100 programs, are critical to keeping people alive, and society and the economy humming, Crowe said. “You could do the entire AHEAD100 project and get all the monoclonal antibodies through phase 1 clinical trials for about US$1–$2 billion dollars. That’s not even a decimal-rounding error for what’s going on right now,” he said, pointing to the estimates of COVID-related U.S. economic losses stretching into the trillions. “So the question is: Now that we’ve experienced this historic pandemic event, will we have the political will to invest $1–$2 billion in interventions ahead of time?”

Uses of antibody therapies
Whether they are RNA- or protein-delivered, antibody therapies could have several applications, according to Denny, Sempowski and Crowe:
• protection for individuals who have not been exposed, notably those at high risk due to health conditions or advanced age;
• post-exposure prophylaxis for those who may have been exposed but aren’t yet diagnosed, such as people identified through contact tracing;
• treatment for infected individuals who have only a low-grade fever, cough, or other mild COVID-19 symptoms, as a way to help stop the disease from progressing and requiring hospital care;
• treatment for infected individuals who are hospitalized with severe COVID-19 symptoms, as a way to help them recover.
Antibodies will not, however, be a replacement for vaccines. “I think you need both,” Crowe said. “If you walked into a room and got exposed, you’d love to have antibodies right now because you might have that virus in your body and you don’t have weeks to wait for vaccine immunity to kick in. But if you want to take a workforce and get them back to work, you want more than temporary antibody coverage: you want something that covers a year or two or more, and that’s going to come from a vaccine.”
Sempowski concurred. Antibodies are a bridge to the longer-term solution of vaccines. “The goal,” he said, “is to get us over the hump until safe vaccines can be approved and made available for mass distribution.”
References
[1] Defense Advanced Research Projects Agency, “DARPA names researchers working to halt outbreaks in 60 days or less,” press release, Feb. 22, 2018. Accessed: Jul. 10, 2020. [Online]. Available: https://www.darpa.mil/news-events/2018-02-22
[2] G. D. Sempowski, K. O. Saunders, P. Acharya, K. J. Wiehe, and Barton F. Haynes, “Pandemic preparedness: Developing vaccines and therapeutic antibodies for COVID-19,” Cell, vol. 181, no. 7, pp. 1458–1463, Jun. 25, 2020. Accessed: Jun. 23, 2020. [Online]. Available: https://doi.org/10.1016/ j.cell.2020.05.041
[3] N. Kose et al., “A lipid-encapsulated mRNA encoding a potently neutralizing human monoclonal antibody protects against chikungunya infection,” Sci. Immunol., vol. 4, no. 35, p. eaaw6647, May 17, 2019.
[4] S. J. Zost et al., “Potently neutralizing and protective human antibodies against SARS-CoV-2,” Nature, vol. 584, no. 7821, pp. 443–449, Jul. 15, 2020, doi: 10.1038/s41586-020-2548-6. Accessed: Jul. 17, 2020. [Online]. Available: https://pubmed.ncbi.nlm.nih. gov/32668443/
[5] S. J. Zost et al., “Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein,” Nature Med., no. 9, Jul. 10, 2020, doi: 10.1038/s41591-020-0998-x. Accessed: Jul. 17, 2020. [Online]. Available: https:// www.nature.com/articles/s41591-020-0998-x
Leslie Mertz (lmertz@nasw.org) is a freelancescience, medical, and technical writer, author, and educator living in Northern Michigan.