In 2011, the California-based company Genomatica reported its success in rigging Escherichia coli microbes to convert sugar into the industrial chemical 1,4-butanediol (BDO). It was a feat of metabolic engineering: BDO is a key ingredient in the production of goods like running shoes, solvents, and spandex. At the time of the company’s announcement, 2.8 billion tons of BDO were produced every year in a multistep, fossil fuel-based process. Genomatica’s system neatly reduced all of that into a cheap, sustainable, one-step fermentation process. The company spent another year refining its technique and finally went commercial with the platform in late 2012. From start to commercialization, the process took about five years.
But then in 2014, a graduate student at the the lab of California Institute of Technology (Caltech) bioengineer Richard tackled the problem; it took her only a few months to produce detectable levels of the same chemical. Soon after, Genomatica senior research scientist Stephanie Culler applied that student’s same technique to the problem from scratch. Pretending, she explains, “that I didn’t know anything about 1,4-BDO, I took all of those pathways that we explored early on, threw them in the system, and, immediately, we beat the titers that the lab team observed after nine months of working on the project.”
What was the difference? The first time Genomatica engineered the microbes, they used traditional molecular biology approaches. Computer models and predictive algorithms helped them decide where to start in terms of basic pathways to try, but actually testing the different systems required laborious cycles of engineering enzymes, cloning DNA plasmids, coaxing that genetic information back into various microbe strains, seeing if it worked, and trying again when it didn’t. This is standard procedure today; and, while it works to an extent, it’s very inefficient. Think of an airplane engineer jumping straight from blueprint to life-size airplane, checking if the airplane flies, and then if it doesn’t, going back, designing a different variation, another airplane, and on and on until the winning iteration eventually emerges.
Murray’s student, however, used a technique developed by a researcher named James Swartz from Stanford University and further advanced by others, particularly Northwestern University and University of Minnesota researchers Michael Jewett and Vincent Noireaux. It was an old, but newly invigorated concept called cell-free biology. Here, the researchers essentially lifted the microbes’ entire DNA-transcription and proteinmaking apparatus out of the cell, put it in dishes, and rapidly ran through permutations of the potential pathways in parallel prototype reactions before translating the more successful combinations back into microbes. There was no working around cell membranes, no plasmid incorporation. They could drop their DNA instructions into a well and see what proteins it produced in the space of eight hours.
Enormous progress has occurred in synthetic biology, but it has come slowly and laboriously. Biologists contend with the field’s still incomplete knowledge about cells and cell pathways, the unwieldiness of working inside organisms—in vivo, that is—and the tug-of-war between what they want from biology and what biology needs to survive. In the wake of a series of technological advances over the past two decades, cell-free platforms have now emerged as a potent alternative. Manufacturing and industrial engineering are actively benefitting from the technique—Genomatica’s success is a perfect example. But now, biomedicine and basic discovery look to similarly benefit, and sophisticated cancer-fighting constructions and synthetic cell organelles are already emerging in research and industry circles.
Within a few weeks of Culler’s initial cell-free experimentation, she and her team successfully established a concrete proof-of-concept showing production of 1,4-BDO and a second chemical called 1,3- BDO, which is used in rubber manufacturing. The company has since begun to apply the approach to their other chemical products, including nylon precursors and additional, so far undisclosed, compounds. “It took years off project timelines,” Culler says.
Under the Hood
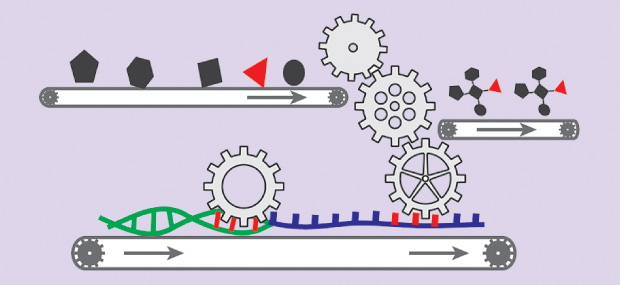
There are two main approaches to cell-free biology. In the first, researchers crack open their cells of choice—usually E. coli—then sift out the unwanted detritus and use the remaining extract as a biological foundation. To that mix, they add their DNA of choice, fuel, and any other elements they believe will help. This is the general approach Genomatica used. A similar alternative relies on purified enzyme systems, wherein the separate elements of a cell’s production engine are picked out, purified, and constituted with the DNA and fuel (Figure 1). It’s a cleaner set-up without the bustle of background reactions that can happen with crude extracts, which makes it ideal for some research purposes, but also comparatively more expensive and less productive.
However, neither of these are new concepts. In fact, two Nobel prizes have already been won using the approach over the last century. But the technique didn’t take off until this past decade, in part because the metabolic systems were so daunting, explains Jewett, a synthetic biologist at Northwestern. “Cell-free biology has been practiced for decades, but for decades, the practitioners have been intimidated somewhat by the complexity of these extracts, which contain hundreds of bioactive catalysts.”
It wasn’t until after a series of studies by Stanford’s Swartz and Dong-Myung Kim (then Swartz’s post-doctoral student) that this began to change. Their work, mostly in cell extracts, showed how these enzyme cocktails could be modified to become more reliable, how adding specific chemicals to the mix could ramp up protein production, and how the use of inexpensive fuels, like amino acids and glucose, could drop the cost of using these systems dramatically—30-fold by 2005. Purified approaches emerged around this time as well, the most prominent of which was developed by Japanese researcher Takuya Ueda at the University of Tokyo, which consisted of a toolbox of purified E. coli enzyme and translational components that could be similarly combined with DNA instructions; this has since been commercialized by several companies, including Cosmo Bio and New England Biolabs.
“There has really been a major shift,” says Jewett, who studied under Swartz during some of those early years. The work showed researchers that these systems of enzymes were something that could be used and harnessed with or without the cell. “This kind of frame shift allowed us to start thinking about and leveraging practices that have been successfully used by chemical engineers and other types of engineers for decades, now applied to these systems of catalysts,” he continues.
Jewett draws an analogy to a mechanic working on a car: “If I want to work on my car, I can basically lift the hood up, see what’s inside, and learn how to take that engine and control it and manipulate it in unique ways. We’re basically doing that with a biological system.”
Powering Production
Genomatica’s prototyping example highlights one advantage to working outside of a cell. In his own lab at Caltech, Richard Murray refers to the idea as a breadboard, as in the basic board that electrical engineers might use to plug in different electrical components, rearranging them and swapping them to test hypothesized design ideas quickly. In biological substrates, the notion becomes a little more abstract, but it’s basically a simpler, external environment, where someone can quickly add different variants, albeit in wells instead of a board. Because there’s no cell, Murray explains, a researcher doesn’t even have to worry about assembling all of the DNA into plasmids. They can literally drop in linear DNA and see what happens. “We can build those variants just by changing the amount and combination of various components we put in,” he says. “We can explore the design space quickly, build things quickly, test them quickly, and then cycle back to design.”
Genomatica elected to return the selected biopathways into cells for final production. Others, however, have discovered that cell-free systems can serve as a production platform in their own right. Here, researchers never have to pit their end goal against the cell’s survival needs. This makes it easier to produce items that might ordinarily be toxic to a cell, like chemotoxic therapeutics. Nontoxic levels of isobutanol have tended to max out at around 2–2.5% v/v in yeast cells, for example; but by using a purified cell-free approach in 2012, researchers from the Technische Universität München, Germany, pushed that yield to 12%. Similarly, Bradley Bundy at Brigham Young University in Utah—another former student of Swartz—reported this year in Biotechnology Journal how his team had been able to use cell extracts to produce the vexingly difficult and highly toxic anticancer drug onconase. Onconase degrades tRNA (an essential component of translation), but Bundy found that spiking the open mixture mid-reaction with extra tRNA boosted yields 56-fold to roughly 2 mg/mL.
“You have this system where you have direct access to the biological machinery, and it’s ripe even for moving to conditions that a cell couldn’t handle—you can change ionic strengths, chaperone concentrations, expression template ratios, and just try to optimize it to the given product that you want,” Bundy explains. “It’s kind of fun because you can mix and match, control the expression yields of different types of proteins, and the ratio of different plasmids.”
It appears that this kind of work can scale up linearly, like a chemical system that works more or less the same way in small batches as in large—as opposed to cell-based systems that can behave differently from one scale to the other. In 2009, cell-free platforms broached industrial levels of production for the first time when the San Francisco, California, company SutroBiopharma increased its production of human cytokine protein to 700 milligrams per liter, in 100-liter vats of E. coli extract.
Sutro is now combining its manufacturing ability with its cellfree technology to design, test, and manufacture engineered antibodies with the ability to bind to specific cancer cells and either deliver a cytotoxic payload or attract the body’s own immune system to wipe them out. With these multicomponent drugs, explains Sutro CEO William Newell, it matters where the different binding elements are and how they’re structured—the shape of the antibody affects its behavior, as does the location of the linker binding it to a drug, the location of that linker on the drug, and so on.
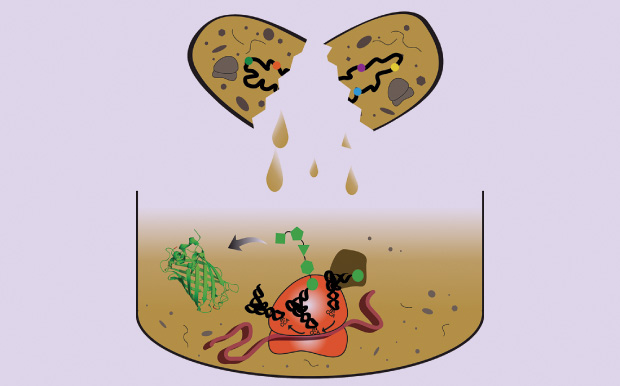
So when scientists design and produce their therapeutics, they’re able to run down the molecular line, mutating every potential amino-acid binding point (Figure 2). “And if you roboticize it as we have, now you can run hundreds to thousands of antibodies generated a day,” Newell says. “Using this empirical method, we can now start to understand how changes in the structural form of the antibody affect the activity.” No products are on the market yet, but the company has completed construction on a manufacturing plant in San Carlos, California, and plans to be ready for inspection by government regulators in 2016.
The potential applications here run far and wide: Boston-based GreenLight Biosciences is industrializing crude extracts similar to Sutro’s to produce a mix of products ranging from food supplements to pesticides, as well as other drugs. Jim Collins, an MIT bioengineer, has experimented with both purified enzyme and extract systems to design a range of programmable cell-free gene networks embedded in paper strips—and crucially unencumbered by cell walls—that could serve as rapid, portable diagnostics for pathogens like ebola and antibiotic-resistant bacteria.
Future Frontiers
Most of the work in this modern cell-free surge has focused on practical biomanufacturing applications. But in the past few years, the technology has also been increasingly used as a platform for research and discovery. For instance, the same qualities that make an in vitro system so appealing for production—speed, toxicity tolerance, and accessibility—also make it a useful vehicle for work expanding the genetic code with nonnatural amino acids. This may allow researchers more freedom in designing proteins with exotic abilities for biomedicine—Sutro’s antibody– drug conjugates use non-natural amino acids—and research into protein structure and function. One particularly potent example of this comes from Hiroaki Suga’s lab at the University of Tokyo, where Suga has designed a flexible synthetic ribozyme-based system—ribozymes are translational tools that act like enzymes—capable of changing almost any amino acid in vitro, generally with only a week’s preparation.
In related areas, researchers like Michael Jewett have begun to use cell-free platforms to modify proteinmaking machinery itself. Two years ago, his team successfully created a one-step method for producing functioning synthetic ribosomes in vitro; earlier this year, they announced their success engineering a “tethered” ribosome, where the two critical subunits of the machine are linked by an RNA chain to ensure that they work together. These successes will also allow researchers to further explore how cellular machinery actually works, to better understand how to harness it for expanding the genetic code in a unique and transformative way.
“The idea is to try to understand what is cell-free production,” says Vincent Noireaux, the University of Minnesota researcher who is trying to harness cell-free extracts as a way to construct such artificial cells for much the same end purpose. He has cited Jewett’s ribosomal work as one of several important milestones in the field. “What is the logic of unicellular cell-free production,” Noireaux asks. “We understand a lot of things in cells, but when we want to put things back together, it does not work. We do not really understand the basic systems.”
Challenges remain in the cell-free field, of course, though researchers are at work to counter them. The basic fuel to power reactions has become cheaper, but other important ingredients in the ingredient mix remain expensive and unstable. Purified systems are ferociously more expensive than crude systems—by roughly 1,000 times in milligram protein produced per dollar in the commercial realm—and their protein yields are generally lower. In lysate work, there’s still no standard way to break open cells for their extracts; so, while high protein yields have been produced in labs such as Sutro’s, this success isn’t universal. At the same time, the advantage of working without cell wall barriers is part of the platform’s appeal, but it also comes with a cost because the systems don’t necessarily have the same ability a cell has to shuffle cellular activities around in different compartments, change concentrations in different parts of the cell, or shunt away bottlenecking intermediaries. Countering these issues, some researchers have begun to experiment with encapsulating parts of a cell-free reaction in artificial membranes, or using microfluidic devices to better recapitulate the cell environment without having to be in the actual cell.
As for the end goal to all of this? “This is all a piece of the broader field of synthetic biology,” says Richard Murray. “What synthetic biology is trying to do is understand what kinds of interesting or useful devices can be created out of biological substrates. We see amazing things around us in nature, so we know that biology and biological components can do amazing things.” Metabolic engineering, artificial cells, and gene circuits that can detect different events are all a part of that, he says, but to progress with them science needs to have a good understanding of what is happening at the biomolecular level and the ability try out different ideas. And cell-free provides another tool with which to do that.



